Determination of the gene regulatory network of a genome-reduced bacterium highlights alternative regulation independent of transcription factors
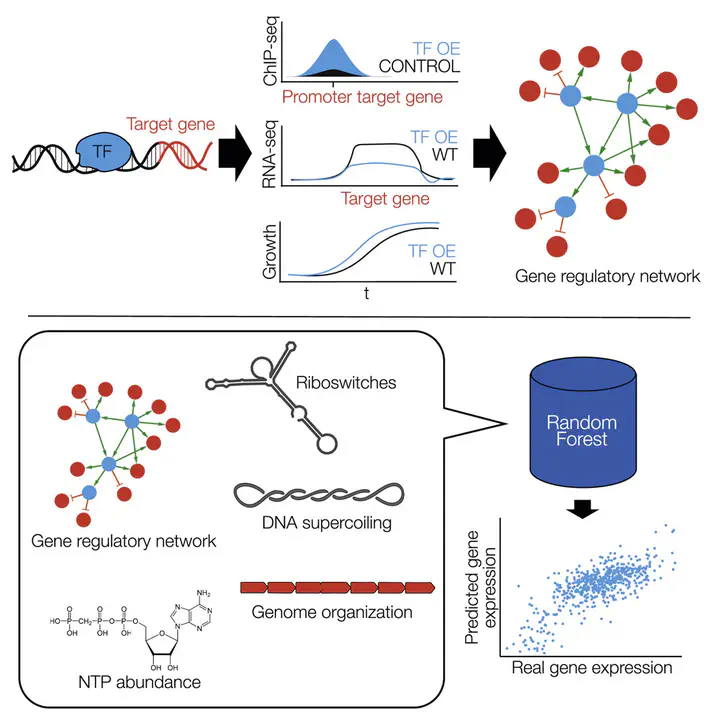 Image credit: Cell Systems
Image credit: Cell Systems
Abstract
Here, we determined the relative importance of different transcriptional mechanisms in the genome-reduced bacterium Mycoplasma pneumoniae, by employing an array of experimental techniques under multiple genetic and environmental perturbations. Of the 143 genes tested (21% of the bacterium’s annotated proteins), only 55% showed an altered phenotype, highlighting the robustness of biological systems. We identified nine transcription factors (TFs) and their targets, representing 43% of the genome, and 16 regulators that indirectly affect transcription. Only 20% of transcriptional regulation is mediated by canonical TFs when responding to perturbations. Using a Random Forest, we quantified the non-redundant contribution of different mechanisms such as supercoiling, metabolic control, RNA degradation, and chromosome topology to transcriptional changes. Model-predicted gene changes correlate well with experimental data in 95% of the tested perturbations, explaining up to 70% of the total variance when also considering noise. This analysis highlights the importance of considering non-TF-mediated regulation when engineering bacteria.